The cytoskeletons (e.g. actin, microtubules, and lamin) are fibrous protein polymers that perform various biological functions within the cell while exhibiting dynamic assembly and disassembly. For example, during cell division, a bipolar structure called the spindle apparatus assembles from microtubules and accurately distributes replicated chromosomes to the newly-created two daughter cells. During interphase, the nucleus, which encapsulates the genomic DNA, is filled with lamin, forming a dense meshwork beneath the nuclear membrane to maintain nuclear integrity and control gene expression by interacting with chromatin. The functions of actin present within the nucleus are also being elucidated, revealing increasingly exciting aspects of the role of the cellular cytoskeleton. In our laboratory, we study the intricate dynamics of cellular cytoskeletons by focusing on eggs and embryos, revealing their roles in genomic stability and dynamics during development.
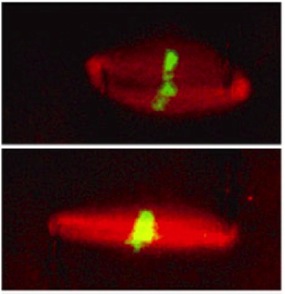
1) Spindle and chromosome segregation mechanicsSuccessful cell division depends on the spindle, a microtubule-based dynamic bipolar machinery. This structure needs to withstand various mechanical stresses that act during cell division while generating sufficient force to segregate chromosomes into two daughter cells. Structural and biochemical properties of the spindle have been extensively studied, but its physical propensities that allow the structure to assemble into a proper bioplar shape and generate relevant forces are largely unknown. We use a combination of biochemistry and biophysics, integrating cell-free Xenopus egg cytoplasmic extracts with force-calibrated microneedles to probe the local mechanical properties of the spindle. The image shows a Xenopus extract spindle (red, microtubule; green, DNA), to which an outward stretching force was applied to near the poles of the spindle to examine the mechanism of force prpagation across the dynamic microtubule-based structure. Spindle dynamics can be monitored using high-resolution spinning-disk confocal microscopy in the laboratory.
References:
|
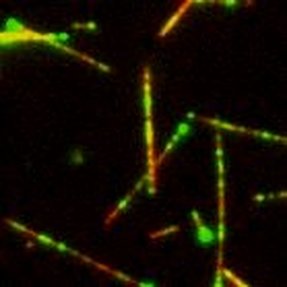
2) In vitro reconstitution of the spindleIn addition to examining the entire spindle micromechanics (see left), we reconstitute a minimal microtubule network in vitro, using a small set of purified spindle components. This allows us to understand how the ordered, large-scale structure of the spindle emerges from interactions of smaller protein 'parts.' Protein concentrations can be controlled using a passivated glass chamber, and microtubule interaction geometry can be manipulated using laser-based nanotweezers. The image shows fluorescently-labeled microtubules (red) and mitotic motor protein kinesin-5 (green) observed using a TIRF-based single-molecule imaging. The microtubule in the middle of the image was mechanically manipulated at sub-micron precision by stirring the laser tweezers to examine how multiple kinesin-5 motors act together to build the spindle.
References:
|
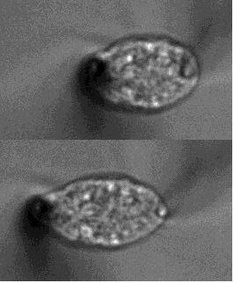
3) Nuclear mechanics and mechano-transductionThe nucleus is a membrane-bound eukaryotic organelle that packages DNA and acts as the site for diverse genomic events, such as transcription. This subcellular structure is subjected to mechanical stresses such as when a cell migrates through a dense extracellular network, contracts in tissues, and attaches to a substrate. Mechanical distortions associated with the stresses are known to damage nuclear structure and perturb the genomic integrity, which are implicated in linking to tumorigenesis and apoptosis. We use microneedle-based quantitative micromanipulation and microrheology methods to examine the material properties of the nucleus and study how the deformability changes depending on lamins and chromatin. The image shows a single HeLa cell nucleus, to which an extensile force is applied using a pair of force-calibrated microneedles. We recently started a new project using mouse early embryos, studying how the nucleus changes its material properties and controls the genome dynamics during development.
References:
|